Information
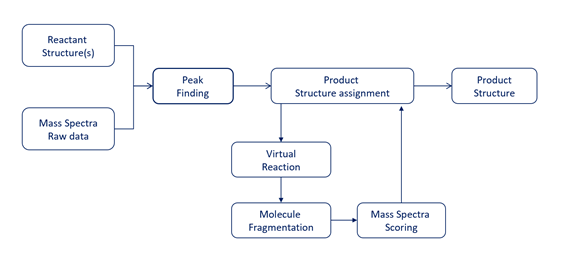
Peak Finding and Structure Elucidation
MassChemSite, first translates the data into a common format standardizing all different types of data input formats. Secondly, the system performs an automatic peak finding using several proprietary algorithms that consider the signal/noise analysis, blank comparison (if provided), compound fragmentation, isotope filtering, mass defect analysis and chromatogram front and tail analysis.
After all these algorithms are applied several chromatographic peaks related to the compound of interest are obtained. These peaks are characterized by their retention time range, m/z observed and the MS2 or secondary spectra. In parallel the system computes the structure of all the chemical products that can be formed applying the chemical rules of the chemical transformations generation a pool of potential structures that are characterized by the structure, their calculated m/z and their potential fragmentation.
The system in the third step assigns the structures to the chromatographic peaks found and assigning a color to each peak:
- Green: This peak color is obtained when the observed m/z can be reached by the product structure calculated m/z computed using the chemical transformation.
- Light blue: This peak color is obtained when the observed m/z can be reached only by computing the selected adducts, dimers or neutral losses to the first or more generation of the product or the reactant.
- Pink: This color is obtained if the peak is only detected in the UV chromatogram and no peak can be assigned using in the MS spectra the UV parameters.
- Orange: This color is obtained if the peak is only detected in the Fluorescence chromatogram and no peak can be assigned in the MS spectra using the Fluorescence parameters.
- Black: This color is obtained if the peak is only detected in the Radio chromatogram and no peak can be assigned in the MS spectra using the Radio parameters.
In the fourth step, each computed structure that is assigned to each peak will be scored in agreement with the fragmentation analysis. This process starts with the fragment assignation made for the reactants by comparing the theoretical fragments obtained by breaking and reorganizing the bonds in the reactant structure and the observed MS and MS2 m/z signals from the spectra. These assigned fragments for the reactant are then compared with the potential fragmentation of the product structure using the same breaking rules used for the reactant.
In this case the fragments found in the products that are shift or non-shifted compared to the reactant ones, or the fragments that are only found in the analysis of the peak spectra and the structure of the product will count as positive in the scoring. The structural fragments of the products that are not compatible with the m/z observed will count as negative in this scoring system. The solutions that have the same scoring will be grouped in a Markush representation.
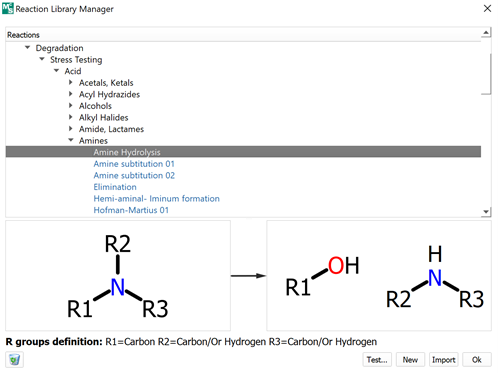
Virtual reaction
MassChemSite includes a module to generate the chemical rules to be applied to the reactants. In this system the user can define a chemical reaction with a high degree of flexibility in the R-group definition. After the rules are defined the chemical pattern will be use in a search first into the structures provided by the user as reactants and it will be applied to generate the chemical structure of the products that then it will be use in the next step as the target to find the chemical rules selected by the user in the reaction definition. The database of reactions can be exported and shared between different user.