MassMetaSite is an established approach for the automatic identification of metabolites for small molecule and peptides using Liquid Chromatography – Mass Spectrometry, UV, fluorescence, and radio-chromatogram data, reducing manual analysis from several hours to only a few minutes per compound. The program can assign chemical structures to each automatically detected chromatographic peak based on the MS and MS/MS fragmentation pattern of the substrates and metabolites.
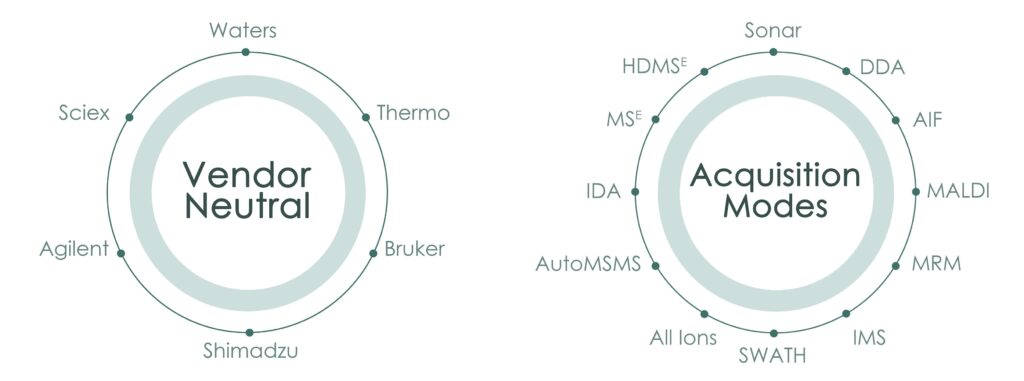
It can process from multiple vendors: Agilent, Bruker, Sciex, Thermo and Waters, and it is also able to analyze data from different acquisition modes: DDA, DDA, MSE, HDMSE, AIF, AF, Broad band, SWATH, Sonar, etc.
In the cases where the data cannot be used to propose a single structure for the chromatographic peaks found, the system also introduces the Site of Metabolism (SoM) prediction from MetaSite computation (the leader in the metabolism prediction market), that ranks the multiple structural options. Moreover, the user has access to the visual analysis of the enzyme-metabolite interaction CYPs, FMO and AOX proteins, and it can even propose structural modification to overcome the metabolic liability.
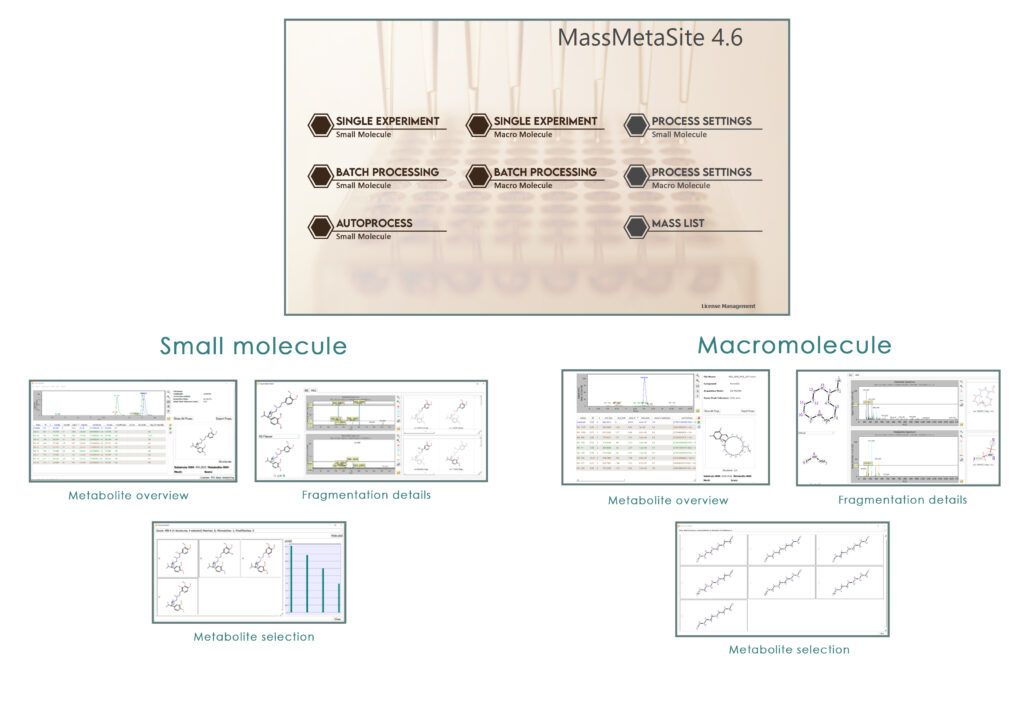
Automatic peak detection
Structure elucidation
Comparison with parent
Metabolite specific fragmentation
Batch processor:
Auto-processing of experiment files
Multiple process batches
New tool to create WebMetabase Protocol Instances/Batches on sample list import time
Exporting to excel report
Markush handling depiction
Modes of operation:
Standard
Standard advanced: user defined mass shift analysis
GSH: Neutral lost and fragment ions
Cyanide
Peptide (up to 4500 amu)
Signal process:
Mass Spectrometry, UV, Fluorescence and Radio files
Direct connection to design tools:
MetaDesign module available, 32D, MetaSite, Metabolite generation, Automatic validation
Isotope labeling:
Stable GSH and cyanide isotope labeling
Radio isotope labeling
Enhanced peak quality analysis:
Area compared to the Blank
Area compared to the Substrate
Isotope similarity score
Improved connection to Waters_connect
Allows missing zero values in Mol or SDF files
Tagging of metabolie names based on custom mass shifts
Custom biotransformations based on formula changes
Automatic update of metabolite formulas based on selected structures in detail view
Vendor Neutral: Waters (.raw, .mzML, waters_connect), Thermo (.raw), Sciex (.wiff,.wiff2), Bruker (.d and tsf), Agilent (.d), SLIM (.d)
Multiple acquisition modes: Waters (MSE, HDMSE, DDA, Sonar, QQQ), Thermo (AIF, DDA, Polarity switching), Sciex (DIA, SWATH acquisition), Bruker (PASEF, TIMSTof, ) and Agilent (AutoMSMS, All Ions)
User defined mass shift peak selection checks
Preferred mass list generation (Waters, Thermo, Agilent)
Possibility to select a UV range for UV chromatogram extraction
Reduced SMS files size
Automatic peak detection
Structure elucidation
Comparison with parent
Metabolite specific fragmentation
Batch processor:
Multiple process batches
Tool to create WebMetabase Protocol Instances/Batches on sample list import time
Visualization in monomers and/or atomic notation
Monomer Database
Charge state from isotopic pattern
Monoisotopic, Most abundant Isotope or average m/z processing methods
Signal process:
Mass Spectrometry, UV, Fluorescence and Radio files
Isotope labeling:
Radio isotope labeling
Enhanced peak quality analysis:
Area compared to the Blank
Area compared to the Substrate
Isotope similarity score
Improved connection to Waters_connect
Allows missing zero values in Mol or SDF files
Read structures as SDF or FASTA format
Process Oligo monomers having a cyclic phosphoamidate group instead of the usual phosphodiester
Better support for oligopeptides
Tagging of metabolie names based on custom mass shifts
Custom biotransformations based on formula changes
Automatic update of metabolite formulas based on selected structures in detail view
Vendor Neutral: Waters (.raw, .mzML, waters_connect), Thermo (.raw), Sciex (.wiff,.wiff2), Bruker (.d and tsf), Agilent (.d), SLIM (.d)
Multiple acquisition modes: Waters (MSE, HDMSE, DDA, Sonar, QQQ), Thermo (AIF, DDA, Polarity switching), Sciex (DIA, SWATH acquisition), Bruker (PASEF, TIMSTof, ) and Agilent (AutoMSMS, All Ions)
User defined mass shift peak selection checks
Improved charge assignment, peak detection and speed for Macromolecules
Added options to expand all monomers
MassMetaSite Training documents
MassMetaSite: Getting started:
Tutorial_MMS_01. Metabolite Identification for Human Liver Microsome Incubation of Verapamil and Data Dependent scan acquisition using Thermo FT Orbitrap
Tutorial_MMS_02. Effect of UV Acquisition: Metabolite Identification for Hepatocytes Incubation of Ebastine and Data Dependent scan acquisition using Thermo FT Orbitrap
Tutorial_MMS_03. Effect of the filtering threshold settings: Metabolite Identification for Human Liver Microsomes incubation of Dextromethorphan in MSe acquisition
Batch Process:
Tutorial_MMS_04. MassMetaSite Batch processing: soft spot batch for h compound. Sample List
Tutorial_MMS_05. MassMetaSite Batch processing: soft spot batch for h compound, directly in the application without Sample List
Tutorial_MMS_06. UNIFI connection. Running samples from a UNIFI server using Ion Mobility data
Reactive Metabolite mode
Tutorial_MMS_07: GSH trapping experiment
Tutorial_MMS_08: Radio labelled experiments
GSH trapping.
Cyanide trapping
Macromolecule:
Single Molecule process:
Tutorial_MMS_09: Analysis of a linear peptide with natural amino acids: the calcitonin case
Tutorial_MMS_10: Analysis of a cyclic peptide with natural amino acids: the somatostatin case
Tutorial_MMS_11: Analysis of a cyclic peptide with natural amino acids: the Somatostatin derivative case
Tutorial_MMS_14: Analysis of an antisense oligonucleotide
Batch process:
Tutorial_MMS_12: Analysis of a family of compounds
Tutorial_MMS_13: Analysis of a time series for Somatostatin
Articles:
Understanding the Metabolism of Proteolysis Targeting Chimeras (PROTACs): The Next Step toward Pharmaceutical Applications
October 2020. Goracci L, Desantis J, Valeri A, Castellani B, Eleuteri M, Cruciani G.J
Metabolite Identification Using An Ion Mobility-Enhanced Data-Independent Acquisition Strategy and Automated Data Processing. Rapid Commun Mass Spectrom
June 2020. Radchenko T; Kochansky CJ; Cancilla M; Wrona MD; Mortishire-Smith RJ; Kirk J; Murray G; Fontaine F; Zamora I
WebMetabase: cleavage sites analysis tool for natural and unnatural substrates from diverse data source
February 2019. Radchenko T; Fontaine F; Morettoni L; Zamora I
Software-aided approach to investigate peptide structure and metabolic susceptibility of amide bonds in peptide drugs based on high resolution mass spectrometry
November 2017. Radchenko T; Brink A; Siegrist Y; Kochansky C; Bateman A; Fontaine F; Morettoni L; Zamora I
Development, optimization and implementation of a centralized metabolic soft spot assay
April 2017. Paiva AA; Klakouski C; Li S; Johnson BM; Shu YZ; Josephs J; Zvyaga T; Zamora I; Shou WZ
Enhancing throughput of glutathione adduct formation studies and structural identification using a software-assisted workflow based on High Resolution Mass Spectrometry (HRMS) data
October 2016. E.N. Cece-Esencan, F. Fontaine, G. Plasencia, M. Teppner, A. Brink, A. Pahler, I. Zamora.
Metabolism study and biological evaluation of bosentan derivatives
October 2016. Lepri S, Goracci L, Valeri A, Cruciani G.
Software-aided cytochrome P450 reaction phenotyping and kinetic analysis in early drug discovery
January 2016. Cece-Esencan EN; Fontaine F; Plasencia G; Teppner M; Brink A; Pähler A; Zamora I
Software-aided structural elucidation in drug discovery
November 2015. Ahlqvist M; Leandersson C; Hayes MA; Zamora I; Thompson RA
Post-acquisition analysis of untargeted accurate mass quadrupole time-of-flight MS(E) data for multiple collision-induced neutral losses and fragment ions of glutathione conjugates
December 2014. Brink A; Fontaine F; Marschmann M; Steinhuber B; Cece EN; Zamora I; Pähler A
High-throughput, computer assisted, specific MetID. A revolution for drug discovery
Spring 2013. Zamora I; Fontaine F; Serra B; Plasencia G
Software automation tools for increased throughput metabolic soft-spot identification in early drug discovery
May 2013. Zelesky V; Schneider R; Janiszewski J; Zamora I; Ferguson J; Troutman M
Update on hydrocodone metabolites in rats and dogs aided with a semi-automatic software for metabolite identification MassMetaSite
May 2013. Li AC; Chovan JP; Yu E; Zamora I
Enhanced metabolite identification with MS(E) and a semi-automated software for structural elucidation
November 2010. Bonn B; Leandersson C; Fontaine F; Zamora I
Posters:
Automation in Metabolite Identification Workflows with Software-Assisted Processing of Mass Spectrometry Data
73rd ASMS Conference on Mass Spectrometry. June 2025. Savannah M Mason1; Ismael Zamora1; Luca Morettoni1; Paula Cifuentes1; Ramon Adalia1
Automatization of structural elucidation workflow for detecting degradation impurities in Peptides
February 2020. Elisabeth Ortega-Carrasco, Blanca Serra, Ismael Zamora
High resolution Mass Spectrometry with automated data analysis to support structural elucidation of degradation impurities of small peptides
AAPS 2019 PHARMSCI 360, San Antonio (United States of America)… 03 November, 2019
Software assisted analysis for Peptide Catabolism
Institute for Research in Biomedicine (IRB Barcelona) 35 EPS European Peptide Symposium, Dublin (Ireland) … August 2018
Peptide catabolite identification using HDMSE Data and MassMetaSite processing
66th ASMS Conference on Mass Spectrometry and Allied Topics, San Diego (United States of America) … 06 June 2018
Peptide metabolism: High resolution mass Spectrometry tool to investigate peptide structure and amine bond metabolic susceptibility
American Peptide Symposium, Whistler (Canada)… 17 June 2017
Kinetic analysis during the metabolite identification process
05 June 2016. 64th ASMS Conference on Mass Spectrometry and Allied Topics, San Antonio, TX (United States of America)
Development of higher-thoughput metabolic soft spot assay with integrated assessment of Glutathione adduct formation
05 June 2016. 64th ASMS Conference on Mass Spectrometry and Allied Topics, San Antonio, TX (United States of America)
Peptide metabolism: Identification of Metabolite structures of GLP-1 receptor agonists in different in-vitro systems using high resolution mass spectrometry
64th ASMS Conference on Mass Spectrometry and Allied Topics, San Antonio, TX (United States of America) … 05 June 2016
Videos:
PROTAC Metabolite Identification
Practical Applications for Drug Discovery 2020 Webinar. Ismael Zamora
Automation of Metabolite Identification
Practical Applications for Drug Discovery 2020 Webinar. Ismael Zamora
Spatial Localization and identification of Drug and Metabolites
Practical Applications for Drug Discovery 2020 Webinar. Sara Tortorella, Giulia Coliva
Metabolite ID For Peptides Made Simple
Practical Applications for Drug Discovery 2020 Webinar. Tatiana Radchenko
Proprietary Peptide Visualization Within MassMetaSite
Practical Applications for Drug Discovery 2020 Webinar. Tatiana Radchenko
Why am I unable to process Thermo RAW files?
Problem
Error messages might be:
RUN ERROR(Unable to process negative control file)
mzqt exception: THERMO INTERFACE ERROR(Unable to init interface)
Unrecognized option: P
Solution
It seems that the computer is missing the Thermo drivers required to work with Thermo files. Specifically, you may not have MSFileReader installed, which is essential for reading these files. You can download MSFileReader from Thermo’s website at the following link:
Download MSFileReader . Please note that you’ll need a Thermo account to access the download. If you don’t already have an account, you can create one by registering for a demo software on their website. Install Thermo MSFileReader in administrator mode.
Install the latest version of the msfile-converter.
Install Visual C++ Redistributable 2012
Install Visual C++ Redistributable 2013
Install Visual C++ Redistributable 2015
Restart MassMetaSite.
Why do I get the message, “error parsing filter line:,” when parsing Thermo files?
Problem
Error message: mzqt exception: THERMO INTERFACE ERROR(error parsing filter line: )
Solution
This error usually occurs due to difficulties accessing the data from network drives. Try again after copying the raw files to a local drive.
Why do I get the 'Unrecognized option: P' when processing Thermo files?
Problem
Error message: ‘Unrecognized option: P’
Solution
Your file-converter is too old for the version of MassMetaSite which is installed on your computer.
Why am I unable to process Agilent .d files?
Problem
Error message: IDISPATCH ERROR (code:Unknown HResult code value: 0x7e CLSID_MassSpecDataReader instance creation)
Solution
Verify that .Net 4 was installed before installing MassMetaSite. To confirm if .Net 4 is properly installed, please check that the folder ‘C:\Windows\Microsoft.NET\Framework64\v4.0.30319’ exists. If not, then .Net 4 needs installed and MassMetaSite needs reinstalled.
Check that Microsoft Visual C++ 2010 x64 Redistributable Package is installed. If it is not, install it.
Check that Microsoft Visual C++ 2013 x64 Redistributable Package is installed. If it is not, install it.
Why am I unable to process Sciex Wiff files?
Problem
Error message might be:
IDISPATCH ERROR
Unable to process negative control file
Unable to get GetSingleSpectrum
Failed to count the number of samples
Solution
If you are unable to get a single spectrum, please check that the .wiff and .wiff.scan files are located in the same folder when the .wiff file is accessed.
If you are unable to count the number of samples, please check that the .wiff and wiff.scan files have read/write permissions. If the files are on a network drive, please try again on a local drive.
If you are unable to process the files or get the IDISPATCH ERROR, the computer may be missing the drivers required to work with Sciex WIFF files (*.WIFF). Please check if the required installer is already in the MassMetaSite installation folder at the following path:
C:\Program Files\Molecular Discovery Ltd\MassMetaSite 4\MMS Wiff Access Patch 2-win64.exe
If the file is present:
Install it by right-clicking on it and selecting ‘Run as administrator’.
If the file is not there:
Download it from the following link: Download MMS Wiff Access Patch 2 (64-bit)
How can I process Waters raw files?
Waters raw files can be read using the file-converter internally installed within the software.
Verify that .Net 4 was installed before installing MassMetaSite. To confirm if .Net 4 is properly installed, please check that the folder ‘C:\Windows\Microsoft.NET\Framework64\v4.0.30319’ exists. If not, then .Net 4 needs installed and MassMetaSite needs reinstalled.
Check that Microsoft Visual C++ 2010 x64 Redistributable Package is installed. If it is not, install it.
Check that Microsoft Visual C++ 2013 x64 Redistributable Package is installed. If it is not, install it.
Where can I find the standard log file in MassMetaSite?
The standard log file in MassMetaSite can be useful for troubleshooting.
Batch processing
In batch processing, the log file is automatically generated for each processing line and stored in the Work Folder. The file name will begin with “log_” as shown below.
You can send this log file to us to help with troubleshooting.
Single experiment processing
In single experiment processing, the log of the current MassMetaSite session can be found in View –> Log Window.
If you prefer to log to a file, then go to Edit –> Preferences.
Check the box to “Log to file” and then provide a location and file name.
Then send the log file to us to help with troubleshooting.
How can I log peak detection in MassMetaSite?
In addition to the standard log file, it can be helpful to have more information about peak detection in order to understand why a peak has not been found by MassMetaSite.
Instructions
Go to Edit –> Preferences
2. Check the box for ‘Log Peak Detection’
3. For batch processing, the standard batch log file will be populated with extra lines for peak detection. For single experiment processing, also check the box to ‘Log to file’ and provide a file name.
4. When the processing is done, compress the log file and send it to us.
Peak detection logging significantly slows down the computation, so don’t forget to uncheck the box when you don’t need it.
Why can’t MassMetaSite find any substrate ions?
Check that the “Retention time range” from the settings does not exclude the substrate peak
Check that the “Ionization mode” is the correct one (e.g., should be negative instead of positive)
Check that enough charge states have been selected to find the expected m/z value
Check that the same peak tolerance is not too tight (e.g. to avoid saturation issues)
Check that the structure in the structure file is correct and compute its mass in a sketcher
Check that substrates imported as macromolecules are not drawn with a formal charge
Check that the signal filtering threshold is not too high (e.g. to find the monoisotopic peak of large peptides)
Try to uncheck the “Pattern filtering” option
Check that the substrate peak is inside the file by opening the vendor software
Why isn’t my parent molecule neutralized when imported in Macromolecule mode?
Indeed this is a known current limitation of the software. Importing the structure in macromolecule mode currently takes the structure as is. Neutralization of the structure is planned in the roadmap though.
Can I add my own biotransformation mechanisms to MassMetaSite?
Users are not able to add new biotransformation mechanisms. However, if there is something specific you think is missing, please contact us as it might be useful for us to add to MassMetaSite. Alternatively, you can add a custom mass shift list.
Mass Shift List: Users can import a file containing biotransformation names and formulas. Shift of masses corresponding to that formula will be identified from the raw data. Change of formula and biotransformation name will be reported. No structure will be assigned to the compound, but some MS/MS information can still be used. Compounds will be colored light blue in the chromatographic plots.
Instructions
Go to the MassMetaSite Process Settings.
Go to the Peak Detection tab à Advanced –> Import file.
3. Two file formats are currently supported:
CSV: A comma separated file where the first column contains the biotransformation name while the second column contains the formula change. No header should be provided
A Compound Discoverer (Thermo Fisher Scientific Inc) exported XML file containing biotransformations
Can I generate a report from the results in MassMetaSite?
Reports can be generated during MassMetaSite batch processing.
Instructions
Go to Edit –> Preferences
2. Check the box to ‘Generate a report file in xlsx format.’
3. The report for each batch line will be generated as an xlsx file and will be stored in the work folder for each batch line.
How many resources do we need to process data in MassMetaSite?
For small molecules, we recommend at least 1 CPU and 3 GB RAM per MassMetaSite calculation.
For macro molecules, we recommend 1 CPU and 8–32 GB RAM per MassMetaSite calculation. For smaller macromolecules, such as macrocycles, we recommend 8 GB RAM. For larger macromolecules, such as insulin, we recommend 16 GB RAM. If you have many reaction mechanisms to consider and much of the structure is expanded (rather than the monomer depiction), then we recommend 32 GB RAM.
For both small and macro molecules, we recommend having additional resources if you would like to run multiple processes in parallel.
Can I export/import Process Settings in MassMetaSite?
Instructions
In the lefthand panel of the Process Settings window, right click the setting of interest. Click Export Setting.
2. Select a location and provide a file name. Click Save. Note: This export method will save the Settings as a .json file. If you need to import the Settings into WebMetabase, they must be saved as .set or .cset.
3. To import the settings, open the Process Settings window. On the bottom left of the window, click Import.
4. Navigate to the location with the saved settings and click Open. The settings are now available in the list in the lefthand panel.
Can I export "Process Settings" from MassMetaSite to be imported into WebMetabase?
Yes. Settings should be exported as .set or .cset files in order to be used in WebMetabase.
Instructions
Open a MassMetaSite experiment which used the desired settings.
In the upper right panel, click ‘Export Props.’
3. Provide a File name and save as a .cset file type.
These settings can now be imported into WebMetabase.














