MetaSite is a computational procedure that predicts metabolic transformations related to cytochrome and flavin-containing monooxygenase mediated reactions in phase I metabolism.
The MetaSite algorithm is unique in being the only software that is not training set dependent, and therefore exhibits improved predictive performance for novel pharmaceutical compounds. MetaSite considers both enzyme-substrate recognition, which is a thermodynamic factor, and the chemical transformations induced by the enzyme, which is a kinetic factor.
The MetaSite flowchart was recently improved by experimental information from the Human CYP Consortium Initiative, a joint venture between pharmaceutical companies and Molecular Discovery, working together to solve the most important issues in drug metabolism (for further information on CYP Consortium Initiative, see this page).
MetaSite gives unprecedented prediction performance, and has now been updated to include the major FMO isoform (FMO3). Key features include the automatic suggestion of fragment modification to optimize specific metabolic issues (MetaDesign), and an interaction map of the substrate with the enzyme cavity (32D) to aid optimisation in the context of the enzyme.
Validation of MetaSite in CYP-Consortium data and in more than forty pharmaceutical companies has shown that the primary site of metabolism was found within the top three MetaSite predictions for more than 90% of the cases.
In addition, MetaSite provides the structure of the metabolites formed with a ranking derived from the site of metabolism predictions. The method also highlights the molecular moieties that help to direct the molecule in the cytochrome cavity such that the site of metabolism is in proximity to the catalytic centre.
Directly blocking the primary site of metabolism can risk creating an inhibitor of the cytochrome, or may negatively affect the activity or selectivity of the compound towards its therapeutic target. Modifying these contributing moieties that most influence the site of metabolism can bypass both of these potential problems.
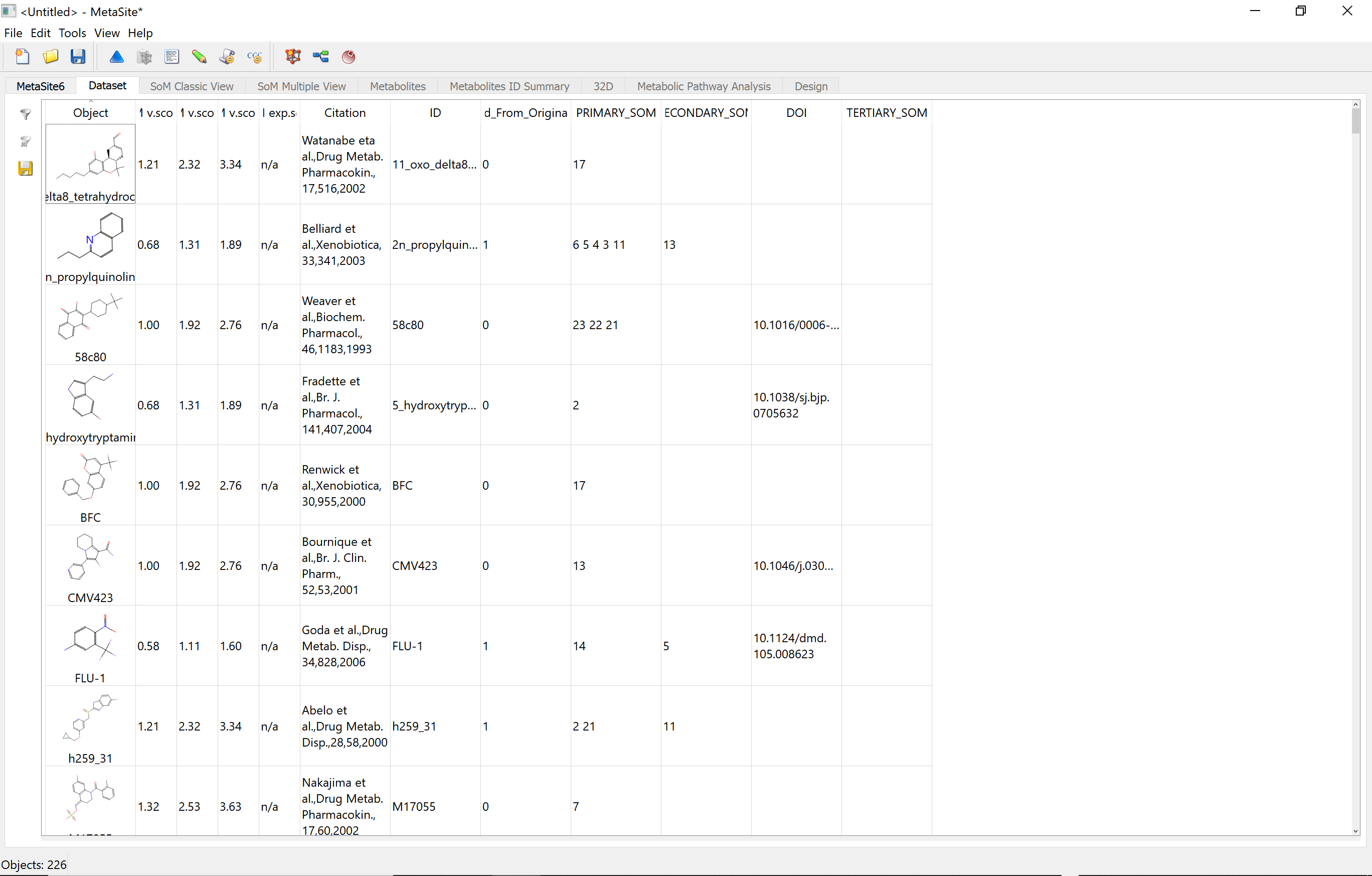
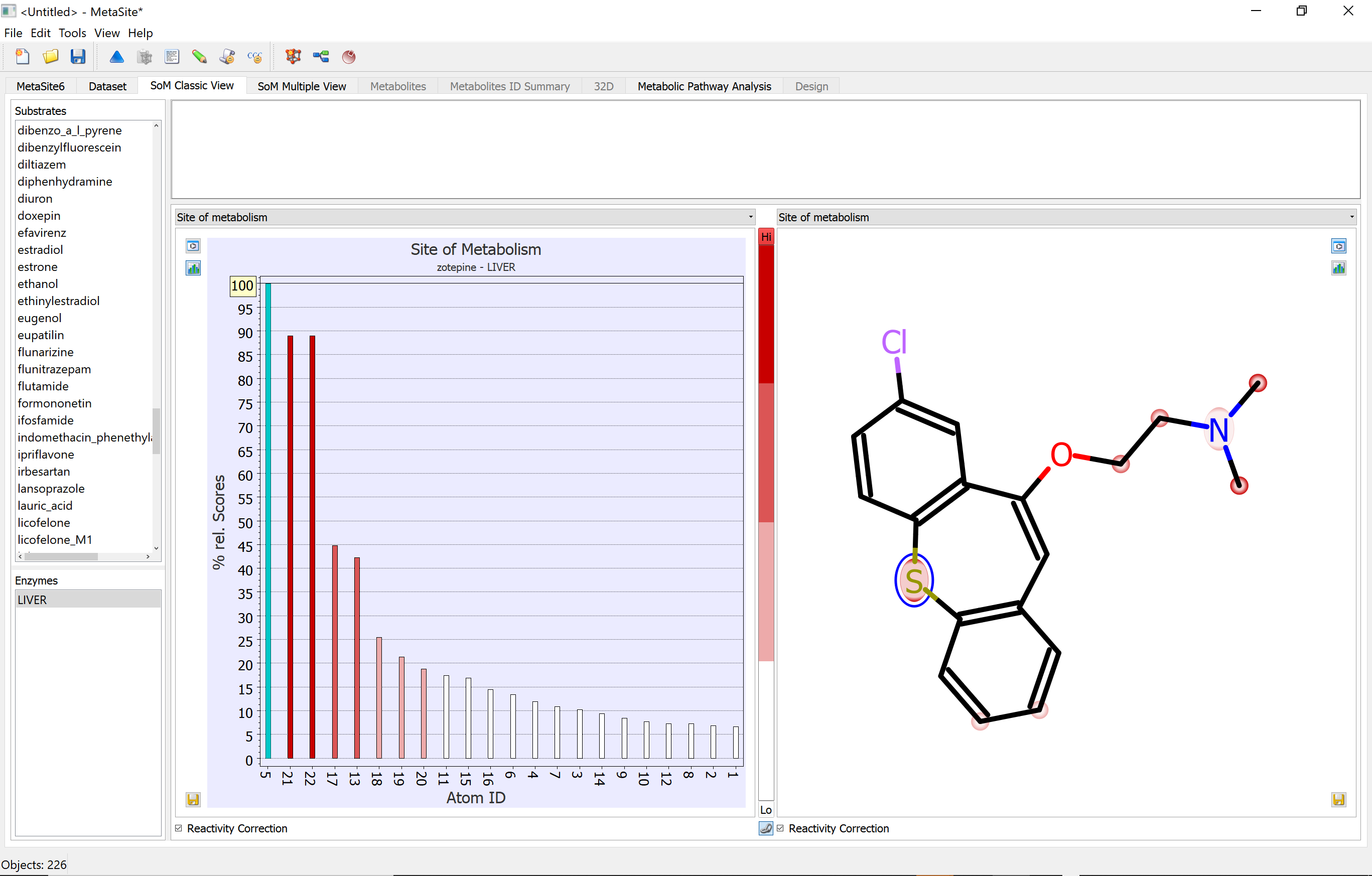
MetaSite Training Documents
- Tutorial_MS_01 Site of Metabolism Prediction
- Tutorial_MS_02 Metabolite identification
- Tutorial_MS_03 MetaSite in Drug Design
Articles:
- Structure-metabolism relationships in human- AOX: Chemical insights from a large database of aza-aromatic and amide compounds
- April 2018.Lepri S, Ceccarelli M, Milani N, Tortorella S, Cucco A, Valeri A, Goracci L, Brink A, Cruciani G
- From Experiments to a Fast Easy-to-Use Computational Methodology to Predict Human Aldehyde Oxidase Selectivity and Metabolic Reactions
- January 2018. Cruciani G, Milani N, Benedetti P, Lepri S, Cesarini L, Baroni M, Spyrakis F, Tortorella S, Mosconi E, Goracci L
- Software-aided cytochrome P450 reaction phenotyping and kinetic analysis in early drug discovery
- January 2016. Cece-Esencan EN; Fontaine F; Plasencia G; Teppner M; Brink A; Pähler A; Zamora I
- Flavin monooxygenase metabolism: why medicinal chemists should matter
- December 2014. Cruciani G, Valeri A, Goracci L, Pellegrino RM, Buonerba F, Baroni M
- Exposition and reactivity optimization to predict sites of metabolism in chemicals
- Spring 2013. Cruciani G, Baroni M, Benedetti P, Goracci L, Fortuna CG
- CYP2C9 Structure−Metabolism Relationships: Optimizing the Metabolic Stability of COX-2 Inhibitors
- August 2007. Ahlström MM, Ridderström M, Zamora I, Luthman K. J
- Comparison of methods for the prediction of the Metabolic sites for CYP3A4 – Mediated metabolic reactions
- June 2006. , , , , and
- MetaSite: Understanding Metabolism in Human Cytochromes from the Perspective of the Chemist
- September 2005. Cruciani G, Carosati E, De Boeck B, Ethirajulu K, Mackie C, Howe T, Vianello R
- Predicting drug metabolism: a site of metabolism prediction tool applied to the cytochrome P450 2C9
- June 2003. Zamora, Ismael; Afzelius, Lovisa; Cruciani, Gabriele














