Specific Workflows
Reactive Metabolite
In addition to the peak finding, structure elucidation and SoM prediction algorithms MassMetaSite offers a specific analysis of the reactive metabolites that may be produced during Glutathione and cyanide trapping experiments or in-vivo situations. The system is looking in one end the neutral losses and the fragment ions that are specific of a GSH adduct formation for both single and double charge species. The value or number of the neutral losses and fragments ions can also be customized.
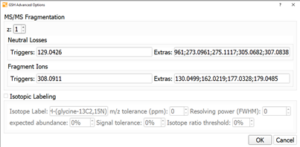
MassMetaSite provides a number of metabolic mechanisms that covers multiple reactive metabolite formation like:
- GSH adduct formation: glutathione over a number of chemical groups (additions to double and triple bonds, substitutions of halides, etc..), also glutathione with double reactions like epoxidation+ GSH conjugation, hydroxylation + GSH conjugation, etc. In addition to the GSH adduct also Cys-Gly and Glu-Cys are considering with the same chemical mechanisms. Also, the reactions with the dansyl derivative of Gluthatione is considered, this chemical is fluorescence compound brings an independent analytical method to detect the adducts. Finally, the system is also ready to use isotope labelled GSH with a custom proportion of labelled and non labelled compound.
- Cyanide adduct formation: The addition of cyanide over iminium and imine is considering, also the double mechanism of formation of iminium over tertiary amine + cyanide conjugation. In addition, the system is also ready to use isotope labelled cyanide with a custom proportion of labelled and non labelled compound.
- Methoxylamine formation: the reaction of aldehydes with methoxetamines is also considered.
Isotope labelling
MassMetaSite also includes a new of algorithms to use isotope filtering and isotope labelling. The user can set up the percentage of hot and cold compound and can set up the expected same peak tolerance in ppm for the isotope as well as the comparison on the difference between the expected and the observed intensity.
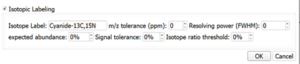
- Isotope Label: In GSH and CN mode, it is the adduct that is used. It cannot be changed by the user. In Radio Labeling mode, it is the radio labeling agent (e.g., 3H or 14C)
- m/z Tolerance (ppm): it is the allowed error between the measured and the expected mass difference (labeled minus unlabeled peak mass).
- Resolving power (FWHM): It is the expected resolving power of the instrument using Full Width at Half Maximum (FWHM) as a reference. It is used to decide if some isotope peaks must be merged (e.g., Chlorine with 14C is not merged at 100,000 but merged at 10,000).
- Signal tolerance: The allowed error between the expected and measured labeled peak height (e.g., Compounds will need at least 70% similarity with the expected peak for tolerance of 30%).
- Expected abundance: The expected signal of the labeled peak compared to the signal of the unlabeled peak (e.g., a value of 200% would mean a labeled/unlabeled 2:1 ratio). In radio-labeled studies, this value is automatically adjusted from the substrate file.
- Isotope ratio threshold: Other isotope peaks having an expected signal value above this threshold will also be considered for the global similarity score (e.g., Chlorine isotope peak will be considered at a value of 10% but not at a value of 40%).
