Chromatography Quality and Multiple signal detection
MS (Mass Spectrometry), Ultraviolet, Radio and Fluorescent
WebMetabase can handle signals from different detector types, performing peak finding in addition to the Mass Spectra chromatogram. The user can select Ultraviolet, Radio chromatography and/or Fluorescence from the analog channel or from external data file sources. All these signals are then analyzed following different peak quality assessment that can also be used to filter chromatographic peaks and automatized using the WebMetabase Macro system.
Peak quality analysis
- Difference between observed and calculated m/z (amu, ppm): For the MS signal the system computes the difference between the observed and the computed m/z. The Observed m/Z considers the m/z finds at the different scans and derived a value which has been compared to the vendor software package to consider effects like peak saturation and loss of accuracy at the top of the peak.
- Comparison to the blank: The system is calculating the ratio between the area of the peak found in the incubation sample under analysis and the black in the case of the MS signal. For Ultraviolet and Fluorescent signal this comparison is done during the peak finding together with a base line correction.
- Comparison to the 0 min sample. In the case that the experiment contains a sample with a 0 min incubation time, the system also does the computation of the ratio between the maximum area found at any incubation condition and the incubation at 0 min. This computation is done for all signal analyzed.
- Area under the curve. In the case that there are multiple incubation time points in the experiment definition, the system performs the calculation of the Area under the curve for each metabolite and for each experimental condition. The number obtained gives an idea of the amount of signal found for each metabolite and it can be particularly useful to compare the “quantity” of the same metabolite at different conditions.
- MassMetaSite scoring, matches and unmatched. Finally, for the Mass Spectra chromatogram for each peak the system also gives the quality of the structure assignment that has been performed by reporting the scoring and the number of fragments that have been found to match in the comparison of the fragmentation schema of the potential metabolite and the parent in the case of the matching fragments, and the number of fragments that goes against the interpretation, in the case of mismatching ones.
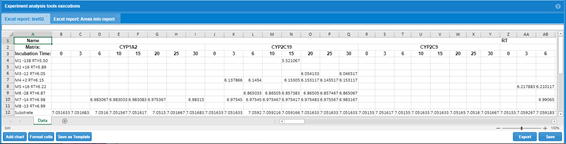
One important consideration with respect to the peak quality measurements is that all this information can eb exported in a Template based report or can directly be output from Chromatogram table in WebMetabase in an Excel report.