Data
Data integrity
WebMetabase is reading the processed data from MassMetaSite or Metabolite Pilot and/or the raw data from external vendor files or UNIFI samples, the system is keeping a copy of the original data so the user can always download and review either the raw or process data.
The WebMetabase workflow for data analysis and reviewing includes two states: in the first one, the experiments are in the Pending state and the user can add/remove/merge chromatographic peaks, change the suggested structure elucidation, modify the fragments that have been used in each structure elucidation, query the raw data, and generate new peaks or re-integrate the existing ones. All the activity will be recorded in the Audit trail called Notebook of the system. Once the user decided to approve the experiment then its status changes to approved and in that situation the user cannot modify the experiment. When the experiments have been approved, it can be used in several analysis tools that can compare experiments across experimental conditions, analysis the impact of different chemical moieties into their metabolism, write reports, etc., but the underlying data cannot be modified. Therefore, the system takes care of the integrity of the original raw and processed data as well as the modifications that the user may introduce. These functionalities together in combination with the user access control to the different modules and functionalities in Oniro help to maintain the system that contains sensitive research and development data.
WebMetabase also provides a system for database consolidation enabling the possibility to integrate data from multiple sources into a single destination. WebMetabase database consolidation allows the user to consolidate some types of data. The system shows to the user a GUI where he can choose the type of data to consolidate and to let him to set the source data (the one to convert) and the destination data (the one it will be consolidated to). The system also allows the user to remove (or not) the source data after its consolidation to the destination data. This possibility is showed only when the customer has the granted permission to remove that type of data. The database consolidation process takes priority over the import process of the experiments, due to database consolidation meaning itself. When a user decides to consolidate a data, he changed all the source data with the destination data in the whole database. So, the system updates the import mapping information with the consolidation information for all the users, independently from which user has done the consolidation.
Audit trail
WebMetabase has an audit trail system called Notebook that records all the operations that are affecting the data, annotating together with the action done, the author (who has made the change in the experiment) and the date. This information is shown in the right panel on every opened experiment, and it can be output to a report, but the user cannot modify it. IF the changes in the experiment are done by the system the user is set to Unknown.
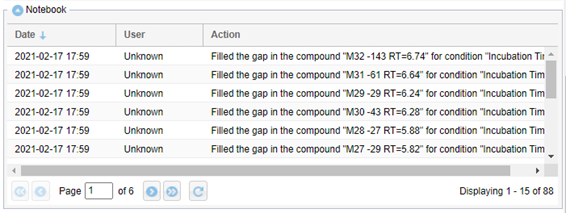
Also, the database consolidation action that is use for data integration is reported in the audit trail of the system. Only users with the enabled role to modify an experiment it can consolidate the data from a particular experiment.

SQL database and Standard data formats
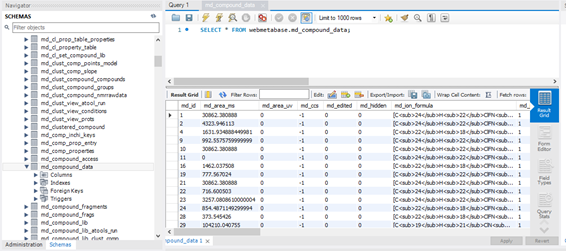
All the information that is produced by WebMetabase it is saved in a SQL database system, that can be under several systems: MySQL and Oracle and the preferred ones. The schema of the database is also available for the user. The data it is saved in standard format using mol format for structure description. In this way this database can also be queried directly from the SQL server. The system uses 2 schemas one of the information about the experiment and another one where the raw datafiles are parked. IN the second one the database is used to park the long files that are used as an entire blob of data.
Licensing and Free/easy access to your data
The licensing system is flexible and can be configured to give you access to the different modules of the application. The license system grants to the user the possibility to import new data to the system, but the user can retrieve any data that has been already introduced into the system without the need of having a valid license. Therefore, the data is always available to the user independently of the license.
WebMetabase provides a ResFul API service to communicate with 3rd party software. (link to the Restful API manual)
RESTful (Representational state transfer) is an architectural style used to design Web services. RESTful principles provide strategies to handle CRUD actions on logical resources using HTTP Requests where methods are mapped as follows:
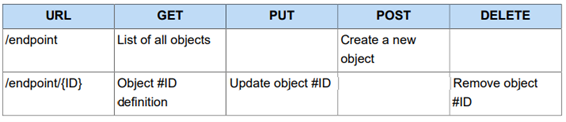
Table 1.1.
Individual resources are identified in requests using URIs (Uniform Resource Identifier)
The base URI for WebMetabase RESTful resources is:
http://SERVER_IP:PORT/WebMetabase
All service requests on resources are done adding the relative resource endpoint to the base URI. For example, to obtain a list of all active users of WebMetabase (where the relative endpoint is /rest/ user), the URI to put in the request is:
http://SERVER_IP:PORT/WebMetabase/rest/user
Most WebMetabase RESTful services can accept requests and produce responses with different MIME types. The Content-type format of the request and the one accepted by the user for the server response are specified in the HTTP Request header. For example, when using Unix command curl, it is possible to specify extra header with the parameter
-H/–header: -H”Content-Type: application/json”
–header “Accept: application/xml”
The Content-type format of the HTTP Response message can also be forced in the request with the URI query parameter “format” (for example format=txt, format=json, format=xml):
http://SERVER_IP:PORT/WebMetabase/endpoint?format=json
The POST and PUT requests on WebMetabase resources return a server response that has the data created or changed by the user request.
