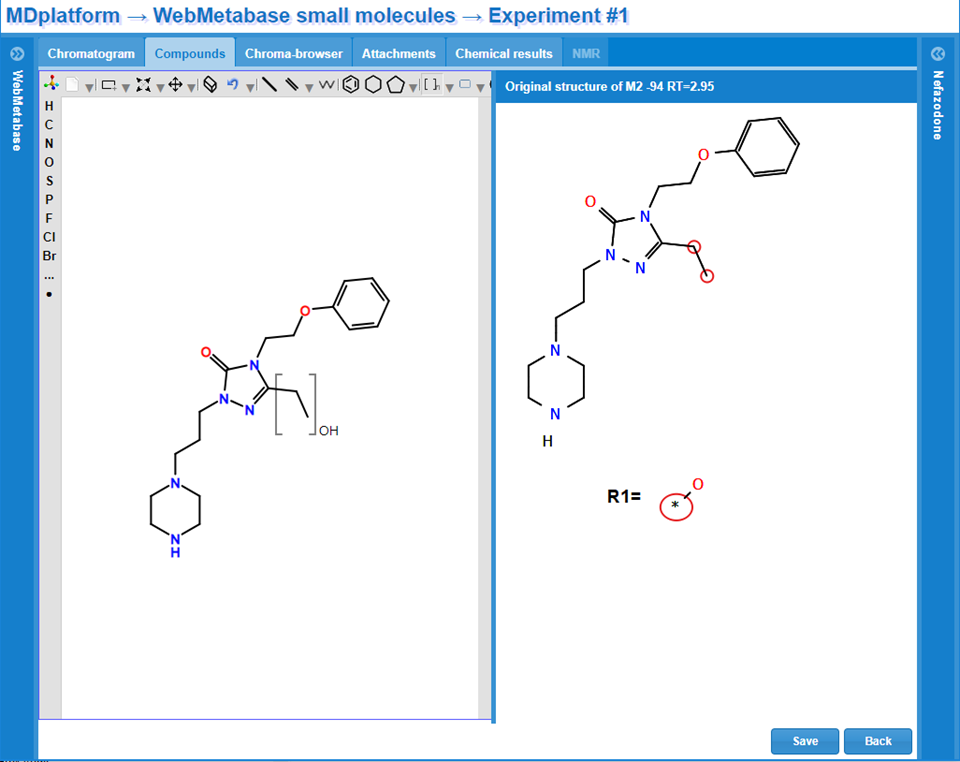
Structure elucidation
Fragment visualization and flexible expert decision tools
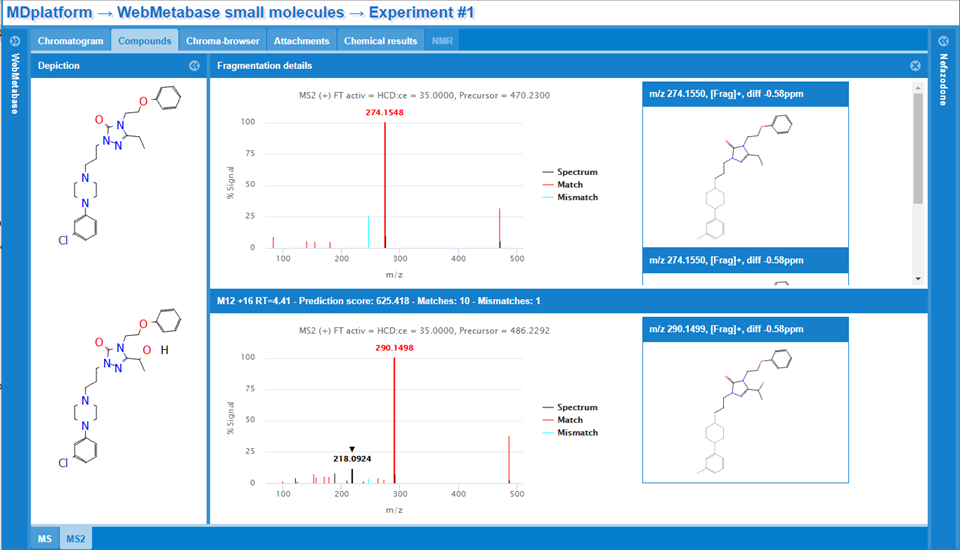
This functionality shows the MS Spectra data interpretation window where the analysis of the fragment structures used to build the Score with the number of matches and mismatches is shown. In that window there are 5 types of fragments that are distinguished by the color in the MS and MSMS spectra:
- Black peaks: There is no fragment assignment to that mass in the parent that can be used in the interpretation of the metabolite under consideration.
- Red peaks: These peaks correspond to the matching and their structural interpretation goes in the direction of the final proposed metabolite structure. By clicking on the red peaks, the structure assigned to the fragment is shown on the right panel.
- Cyan peaks: These peaks correspond to the mismatching and their structural interpretation goes against of the final proposed metabolite structure.
- Coral Peaks: are metabolite matching peaks, their structural information goes in the direction of the final proposed metabolite structure, but they do not have a substrate fragment matching to them. Those peaks could come from manual edition or form MassMetaSite if metabolite fragmentation is selected in the settings.
- Light Green Peaks: are metabolite mismatching peaks, begin the structural information provided for the peak contrary to the proposed structure under study. Those peaks have no substrate peaks related to them and come from the propagation of a manually edited peak.
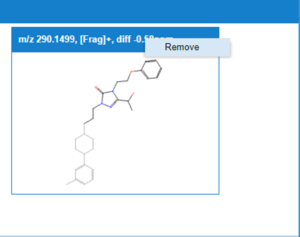
The expert user can decide multiple actions to be considered for each of the assigned and non-assigned peaks:
- Remove peak structural assignment: The suer can select a metabolite peak having a Match or Mismatch assignation and by right clicking on the bar for the metabolite decide to remove it from the analysis. The assignment for the peak is removed for all the proposed structures of the metabolite, and therefore the scores are re-calculated. It is possible that after removing the assignment to the peak, the structure, or structures with higher score change.
Add peak structural assignment: The user by clicking on a metabolite peak that have not assigned into any structural information (black peaks) can draw by themself the assigned structure. The Fragment structure editor allows you to define the fragment by deleting atoms of the parent structure. It will check that the drawn fragment mass is in accordance with the selected peak on the right-hand side panel.
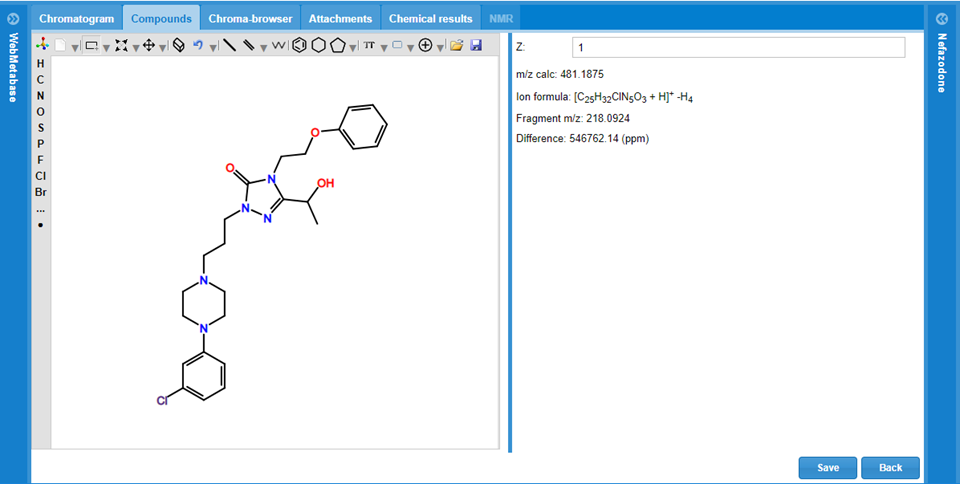
Flexibility to edit the proposed structure
The user has also the flexibility to remove the automatic MassMetaSite actual structural interpretation for the selected chromatographic peak and manually add the user interpretation. All the fragmentation analysis will be removed. The Manual structure editor system is composed of 3 main panels, and the user can draw a single molecule of a Markush:
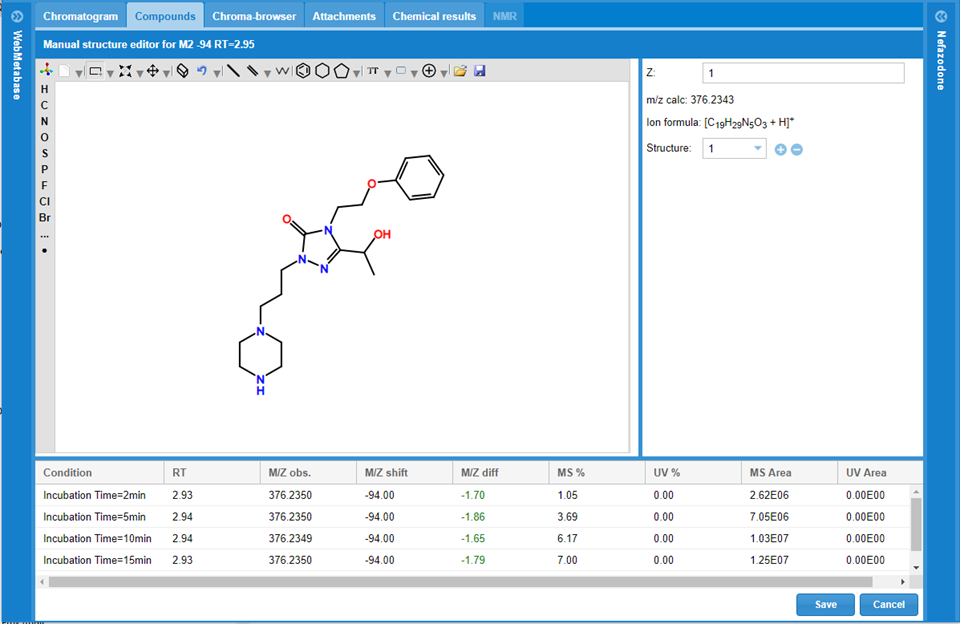
Structure editor: In this panel the user can draw the potential structure associated to the chromatographic peak.
Structure information: Next to structure editor there is the charge declaration for the peak: Z, there the user can change the charge state to use in the calculated m/z which is updated too while editing structure.
In the next line there are the tools to define more than one structure to build a Markush for the new metabolite: pushing the + icon a new structure could be added, and by clicking the – sign a structure will be removed.
Unique metabolite identifier
The user can check if the proposed structure has already been assigned in the WebMetabase database system. When this is done there are 3 options:
- The Metabolite structure is not present in the database. The system search into the data base if there is another metabolite with the same exact Markush structure. In the case that there is not a complete match the system shows the following window, in the left column is the name of the metabolite, in the middle one the Markush representation of the metabolite under consideration and on the right the MS2 spectra.
- It could happen that several structures that compose this Markush could be part of another Markush structure, and therefore by clicking the Other Matches button, the user can have access to those solution for their inspection, although the metabolite ID cannot be assigned to the new one, since the structures are not the same. The structure of the metabolite is shown on the left column, the information about the experiment is shown in the middle column, including the link to the database experiment and the MS2 spectra is shown on the right column.
- The metabolite structure is present in the database (exact Markush match). In this case the metabolite of the other experiment(s) will be shown. The structure of the metabolite is shown on the left column, the information about the experiment is shown in the middle column, including the link to the database experiment and the MS2 spectra is shown on the right column.
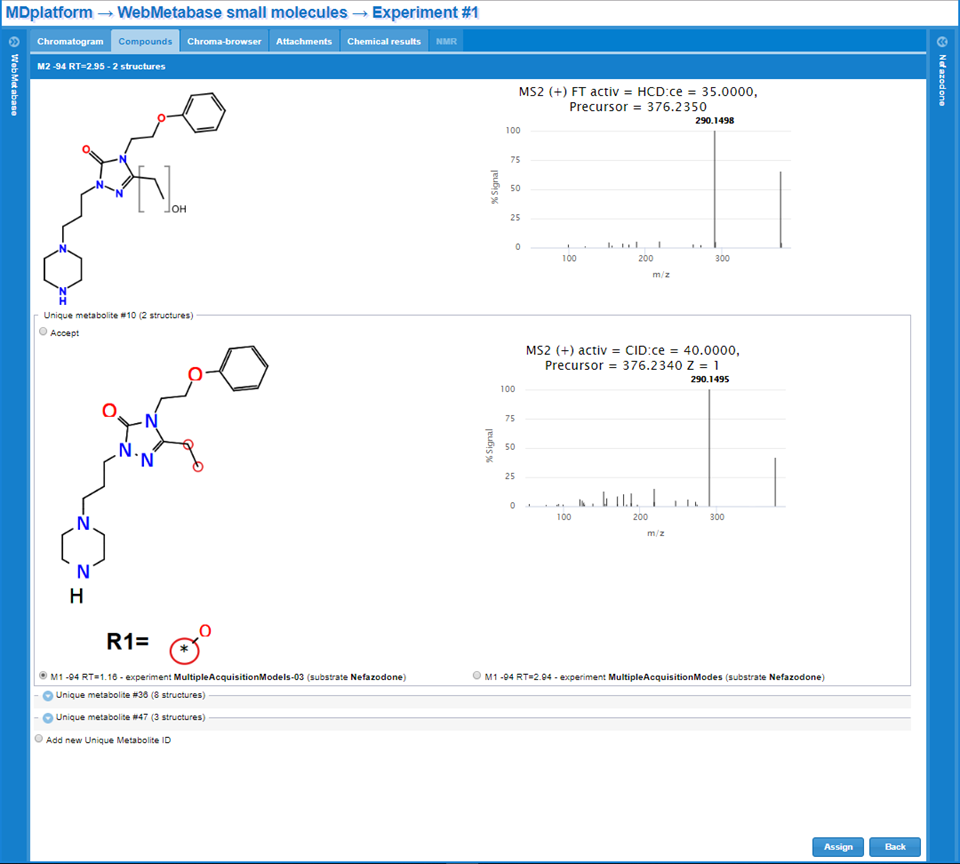
Markush manual drawing
This option is used to replace the automatic “Markush” representation by a user customized depiction, at every place where a “Markush” is represented the custom depicted one will be shown. By clicking this option, a new window will pop-up.