Tutorial_AIQ_02
Tutorial_AIQ_01
Introduction and General Procedure
LC-MS and High-Throughput Data Processing Solutions for Lipid Metabolic Tracing Using Bioorthogonal Click Chemistry
LC-MS and High-Throughput Data Processing Solutions for Lipid Metabolic Tracing Using Bioorthogonal Click Chemistry
24 April 2025
Palina Nepachalovich, Stefano Bonciarelli, Gabriele Lombardi Bendoula, Jenny Desantis, Michela Eleuteri, Christoph Thiele, Laura Goracci, Maria Fedorova
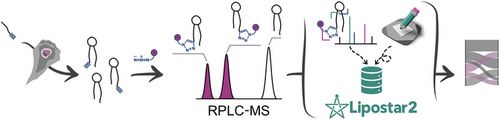
Graphical Abstract
This study introduces an integrated analytical and bioinformatics platform for high-throughput tracing of lipid metabolism using bioorthogonal alkyne fatty acids and optimized LC-MS workflow. Applied to human fibrosarcoma cells, the method traced fatty acid metabolism, revealing nuances in sphingolipid routing and metabolic bottlenecks, highlighting its potential for lipidomics research.
Abstract
Tracing lipid metabolism in mammalian cells presents a significant technological challenge due to the vast structural diversity of lipids involved in multiple metabolic routes. Bioorthogonal approaches based on click chemistry have revolutionized analytical performance in lipid tracing. When adapted for mass spectrometry (MS), they enable highly specific and sensitive analyses of lipid transformations at the lipidome scale. Here, we advance this approach by integrating liquid chromatography (LC) prior to MS detection and developing a software-assisted workflow for high-throughput data processing. LC separation resolved labeled and unmodified lipids, enabling qualitative and quantitative analysis of both lipidome fractions, as well as isomeric lipid species. Using synthetic standards and endogenously produced alkyne lipids, we characterized LC-MS behavior, including preferential adduct formation and the extent of in-source fragmentation. Specific fragmentation rules, derived from tandem MS experiments for 23 lipid subclasses, were implemented in Lipostar2 software for high-throughput annotation and quantification of labeled lipids. Applying this platform, we traced metabolic pathways of palmitic and oleic acid alkynes, revealing distinct lipid incorporation patterns and metabolic bottlenecks. Altogether, here we provide an integrated analytical and bioinformatics platform for high-throughput tracing of lipid metabolism using LC-MS workflow.
